|
4/9/2023 0 Comments Identifies sequences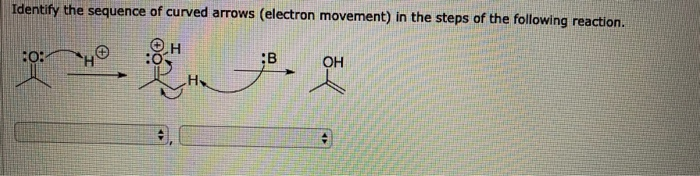 
As clinical assessments/diagnosis of a patient are typically captured in the Condition resource or the ClinicalImpression resource, the MolecularSequence resource can be referenced by the Condition resource to provide specific genetic data to support assertions. The MolecularSequence resource will be referenced by Observation to provide variant information. Hence data such as precise read of DNA sequences and sequence alignment are not included such data are nonetheless accessible through references to GA4GH (Global Alliance for Genomics and Health) API. Implementers should be aware that semantic equivalency of results of genetic variants cannot be guaranteed unless there is an agreed upon standard between sending and receiving systems.įocus of the resource is to provide sequencing alignment data immediately relevant to what the interpretation on clinical decision-making originates from. However, these data should not be used to dynamically correct/change variant representations for clinical use outside of the laboratory, due to insufficient information. discovery research, outcomes analysis, and public health reporting) may use this information to normalize variants over time or across sources. It is strongly encouraged to provide all available information in this resource for any reported variants, because receiving systems (e.g. Tissue source used to determine genotype of the variation.This resource is designed to describe sequence variations with clinical significance with information such as: Reference Sequences: RefSeq and ENSEMBL.Variant Databases: dbSNP, ClinVar, and COSMIC.Thus, the FHIR MolecularSequence resource avoids large genomic payloads in a manner analogous to how the FHIR ImagingStudy resource references large images maintained in other systems.įor use cases, details on how this resource interact with other Clinical Genomics resources or profiles, please refer to implementation guidance document here.ġ0.6.1.1 Genetic Standards and Resources include: Complete genetic sequence information, of which specific genetic variations are a part, is reported by reference to the GA4GH repository. By this method, a sequence can be reported. Atomic sequences can be connected by link element and they will lead to sequence graph. The MolecularSequence resource is designed to describe an atomic sequence which contains the alignment sequencing test result and multiple variations. The committee is considering options for modifying the resource and anticipates potential changes being brought forward in an upcoming ballot. The Clinical Genomics committee has identified overlaps and redundancies between content in the MolecularSequence resource and content in Observation profiles in the evolving Implementation Guide for Clinical Genomics Reporting found here.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
